The BNMA BN Repository
This repository is a resource for posting and downloading Bayesian network models for sharing with others and for providing supporting material for publications. Please respect authors' rights where noted.
Search
5 BNs found.
Predicting Age of Martens From Capture Data
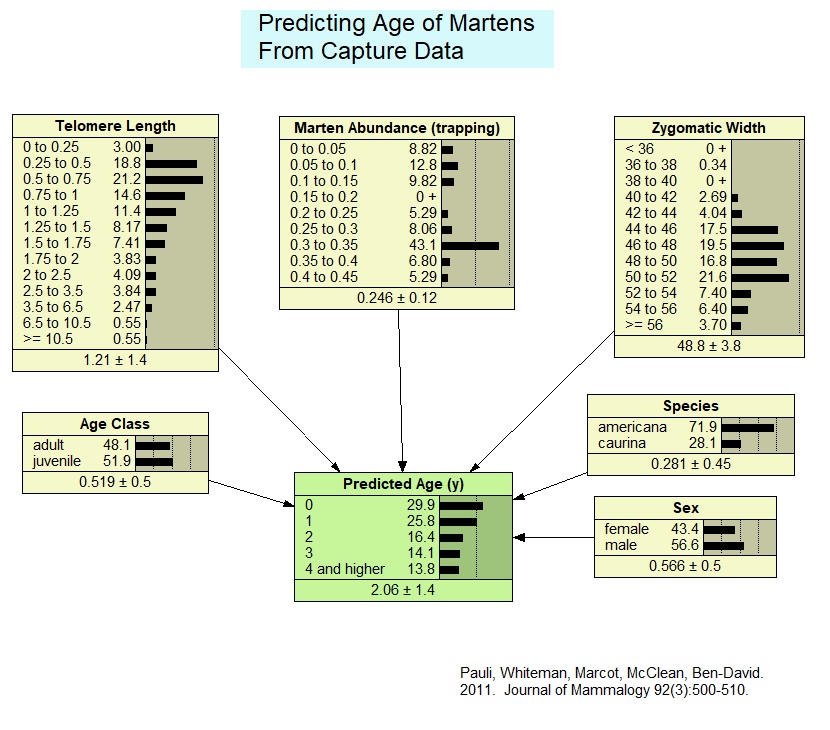
This model derives from a published version, slightly reformatted for testing an application of a Generative Adversarial Bayesian Network (GABN) framework. The model predicts the age of individual martens (members of the weasel family) based on six predictor attributes of marten sex, subspecies designation, zygomatic width (a measure of jawbone size), marten population density (derived from field trapping), telomere length (ends of the chromosomes that shorten over an individuals' life), and general age class (juvenile or adult). The model was initially parameterized with a case file of 399 samples of martens, and then used to generate a fictitious case file using Netica's "simulate cases" function, for testing the GABN approach.
Animals

With this network (also available from <www.norsys.com...>) you can enter some characteristics of a particular animal, and watch how the probabilities of its other characteristics (and what type of animal it is) change.
This is just a toy example. For a real-world application, it would have to be extended to include many animals (or plants, bacteria, etc.), probably all from the same environment, or the same subclass, etc. Also, the "Animal" node should probably have an "Other" state.
The fun part of this network is to extend it to include more animals and more characteristics. You may need to define other groupings, such as the "Class" node, in order to keep things manageable. If you make a great network, send it to Norsys; we would love to include it in our library (with the proper credits).
Marten Telomere Genetics for Age Determination

These models, trained from laboratory and field data, predict the age of individuals of two species of marten (Martes americana and M. caurina) in North America, from relative length of telomeres (fragmented ends of chromosomes that shorten over time) and other anatomical and environmental factors. It is the first time that age of an organism can be predicted from telomeres (and other factors) in a reliable manner. Two models are presented here: the "live capture" versions, for predicting age in years from captured (or museum specimen) martens; and the "noninvasive" version for predicting age class from non-invasive genetic samples where the actual animal does not need to be captured and handled. A case file for running the live capture model version can be found at <abnms.org...>.
Pacific Walrus
We developed a Bayesian network model to integrate potential effects of changing environmental conditions and anthropogenic stressors on the future status of the Pacific walrus population in the Chukchi and Bering Seas, at four periods through the twenty-first century. The model framework allowed for inclusion of various sources and levels of knowledge, and representation of structural and parameter uncertainties. Walrus outcome probabilities through the century reflected a clear trend of worsening conditions for the subspecies.
Recreation Effects on Brown Bears
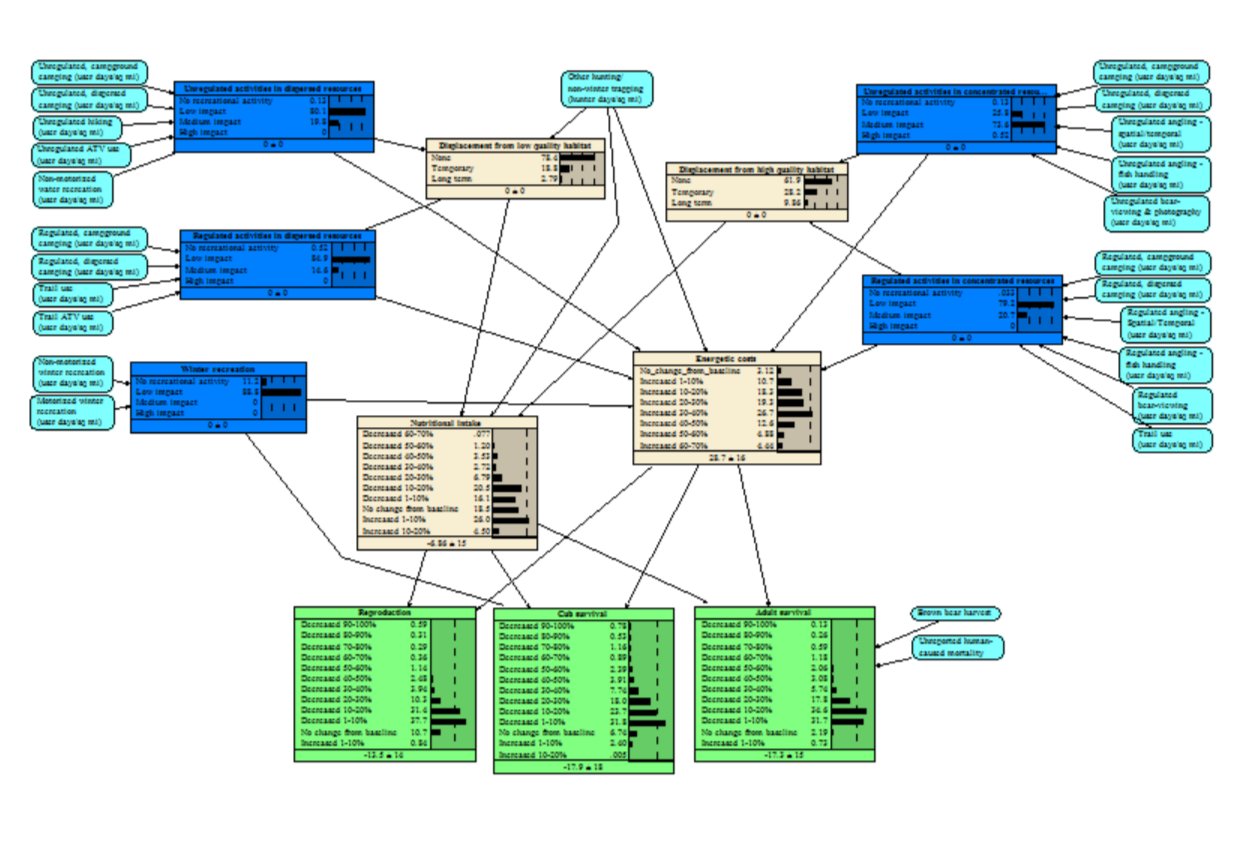
We integrated results from empirical studies with expert opinion to better understand the potential population-level effects of recreational activities on brown bears. We conducted a literature review and Delphi survey of brown bear experts to better understand the frequencies and types of recreations occurring in bear habitats and their potential effects, and to identify management solutions and research needs. We then developed a Bayesian network model that allows managers to estimate the potential effects of recreational management decisions in bear habitats.
 Bayesian Intelligence
Bayesian Intelligence